-
Hit to Lead
- QSAR
- Molecular Dynamics Simulation
- Relative/Absolute Free Energy Calculation
Among the VS approaches, quantitative structure–activity relationship (QSAR) analysis is the most powerful method due to its high and fast throughput and good hit rate. For QSAR model development, Binary star has well-established relevant chemogenomics data collected from databases and the literature. Our established machine learning techniques proved successful in identifying perspective compounds with desired properties.

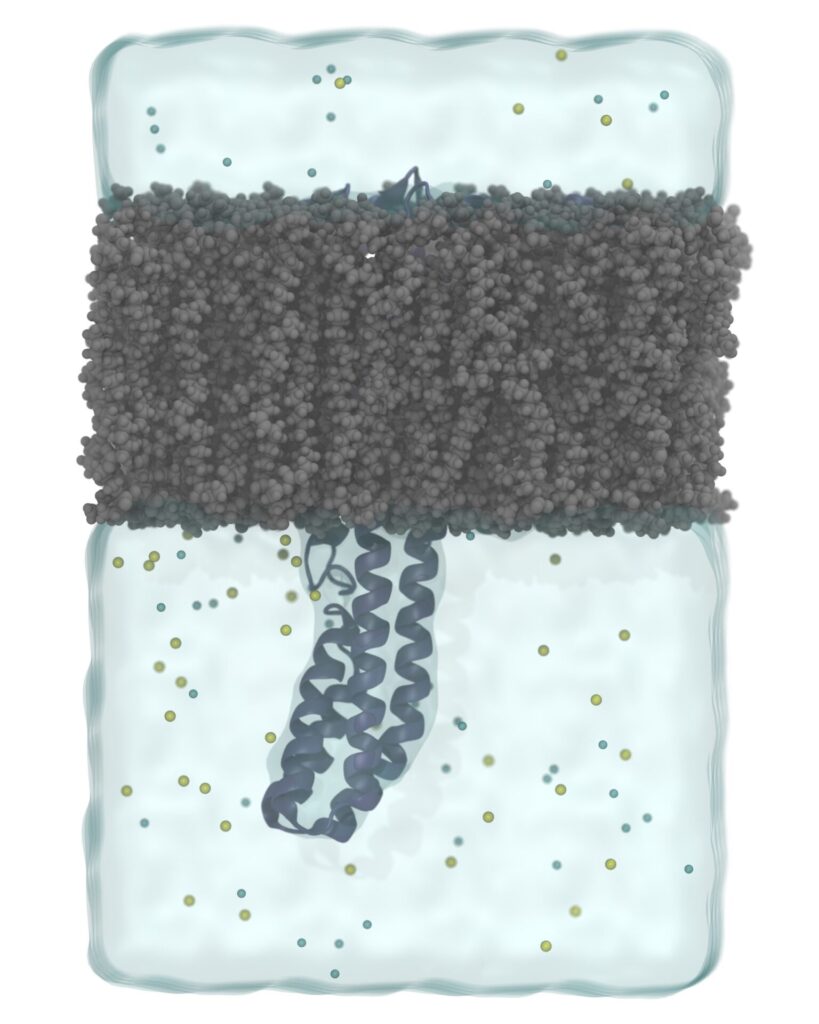
MD simulations describe the movement, interactions, and dynamics of biomolecules (e.g., proteins and nucleic acids) at the atomic level. MD simulations employ a “force field” based methodology that reflect all interatomic interactions and analyze the time dependent behaviour of biomolecules using Newton’s laws of motion. Combining MD with other CADD based techniques allows researchers to have a more realistic approximation of the biological systems. Our molecular dynamics simulation services include conventional long-run (> 1 µs) all-atom and coarse-grained MD simulations. For specific applications, we further employ enhanced-sampling techniques such steered molecular dynamics, adaptive biasing force, replica exchange MD, gaussian-accelerated MD and constant-pH MD simulations. In addition, we run allostery and druggability simulations to predict druggable binding sites of various protein targets by employing long-run mixed-solvent MD simulations.
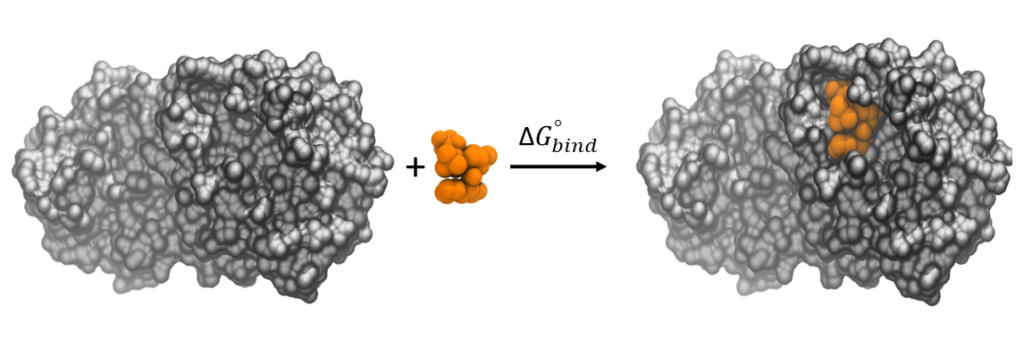
The accurate prediction of ligand-protein binding free energies is crucial in hit identification and hit-to-lead optimization. We run rigorous absolute binding free energy calculations using different protocols such as the alchemical free energy perturbation (FEP) method and the geometrical approach using the adaptive biasing force (ABF) method. We also perform end-point relative binding free energy calculations using various protocols such as molecular mechanics Poisson–Boltzmann surface area (MM/PBSA) and molecular mechanics generalized Born surface area (MM/GBSA) and relative-FEP method.